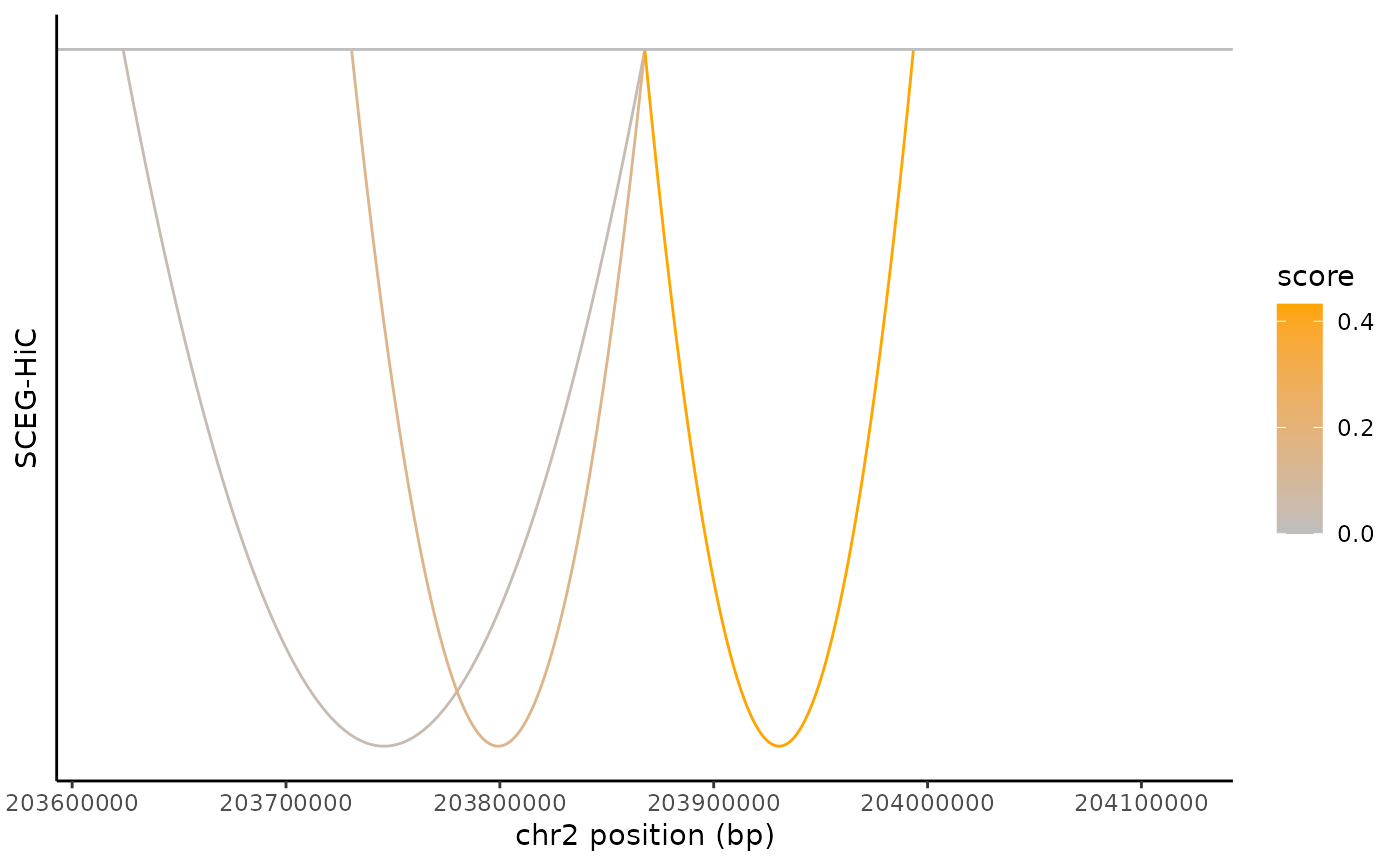
Visualize predicted links between genomic elements (e.g., genes and regulatory peaks) within a specified genomic region.
Arguments
- links
A data.frame containing enhancer-gene links.
- gene
A character string specifying the target gene to highlight.
- region
A
GRangesobject specifying the genomic region to plot.- lowcolor
Color for the lowest scores in the gradient.
- highcolor
Color for the highest scores in the gradient.
- titlename
A character string specifying the title of the plot.
- gene_anno
A
data.frameorNULL. If provided, should be compatible with theGeneRegionTrack-class. Column names must not containNA.
Value
A ggplot object visualizing enhancer-gene links.
Examples
links <- data.frame(
gene = "CTLA4", peak = c("chr2_203623664_203623982", "chr2_203730093_203731243", "chr2_203992800_203993979"),
score = c(0.03623216, 0.14814205, 0.43240254)
)
library(GenomicRanges)
region <- GRanges(seqnames = Rle("chr2"), ranges = IRanges(start = 203617771, end = 204117772))
geneanno <- annotateTSS("Homo sapiens", "hg38")
colnames(geneanno) <- c("chr", "gene", "tss")
links <- LinksPlot(links, geneanno, gene = "CTLA4", region, lowcolor = "blue", highcolor = "orange", titlename = "SCEG-HiC")
links